Service Track 01 | CADD | Foundations of Bioinformatics, Structural Analysis, and Molecular Docking Services

About Bundle
Service Track 01 is a structured, service-oriented training bundle designed for learners, researchers, and early-stage scientific professionals who want to build a strong practical foundation in bioinformatics, protein analysis, and molecular docking, then turn that knowledge into a clear, usable, and professionally relevant analysis service list.
This is not a full FAB Program, and it is not just a collection of unrelated courses. It is a focused Service Track built to move the learner from scientific understanding into analysis-ready execution in a specific and valuable part of the Computer-Aided Drug Design (CADD) workflow. Instead of trying to cover every possible business and freelance layer, this track concentrates on one important goal: helping the learner become capable of performing a clearly defined set of scientific analyses that can later be presented as professional services.
The bundle starts with a strong foundation in Chemoinformatics and Computational Drug Discovery, introducing the learner to the logic of CADD, molecular modeling, molecular docking, virtual screening, molecular dynamics, and scientific workflow design. It then builds the data backbone of the learner’s skill set through chemical databases, protein databases, and the molecular file formats used in real computational drug discovery pipelines. This includes hands-on familiarity with platforms such as PubChem, DrugBank, ZINC, ChEMBL, PDB, UniProt, InterPro, and KEGG, as well as formats such as SDF, MOL, and PDB.
After that, the track moves deeply into protein-centered analysis. Learners study protein structure and function, secondary and tertiary structure understanding, folding and misfolding logic, protein modeling principles, homology modeling, ab initio prediction, protein visualization, structural validation, and model quality assessment. This means the learner does not only see proteins as biological concepts, but as computational objects that can be modeled, analyzed, evaluated, and used in downstream research and drug discovery workflows.
The next stage focuses on protein sequence analysis and evolutionary interpretation. The learner is trained in pairwise sequence alignment, multiple sequence alignment (MSA), conserved region analysis, and phylogenetic tree construction and interpretation. This adds an important comparative and functional layer to the workflow, helping the learner understand how sequence-level information can support structural interpretation, target understanding, and broader research logic.
The track then expands into gene prediction and protein–protein interaction (PPI) analysis, allowing the learner to explore how genomic and proteomic data can be translated into practical analytical outputs. Through this layer, the learner begins to understand not only isolated protein targets, but also interaction networks, biological context, and computational interpretations relevant to research and systems-level thinking.
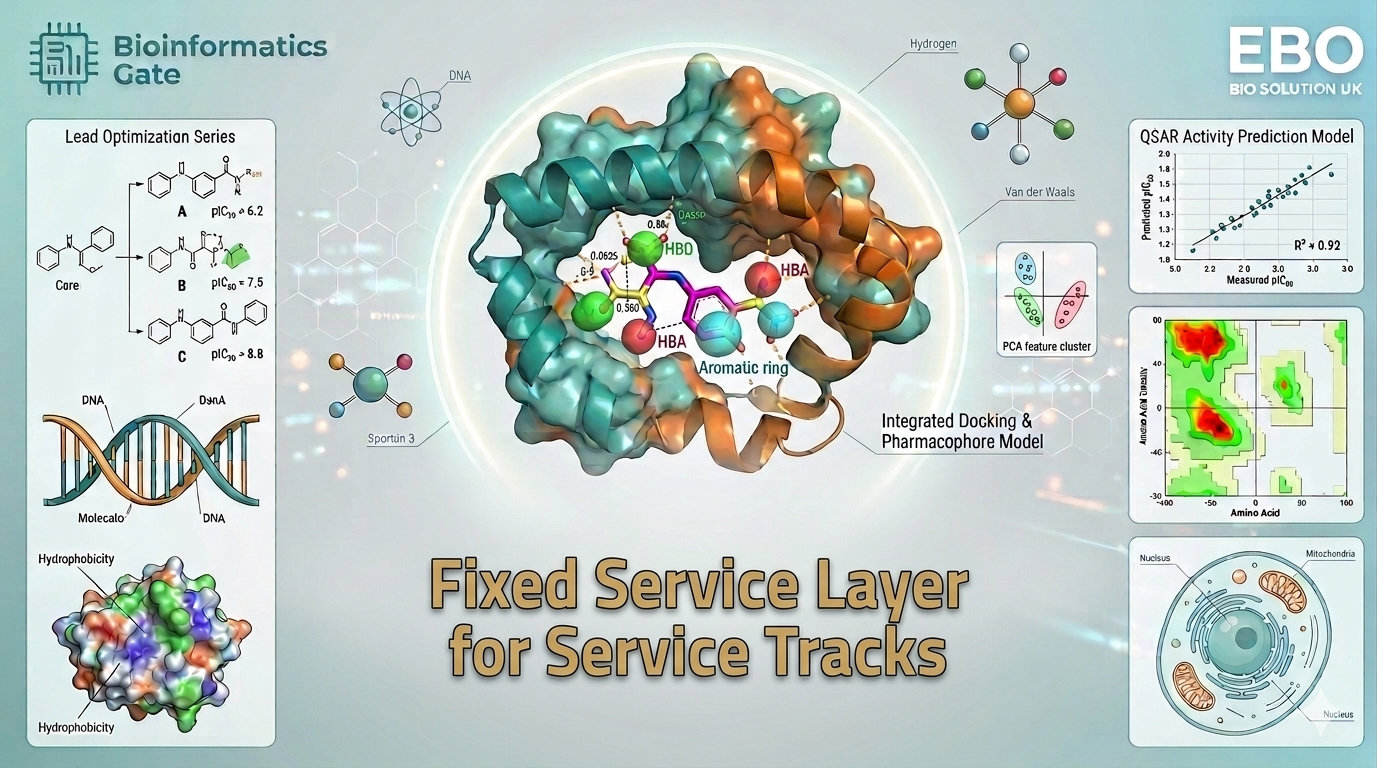
Once that foundation is built, the learner enters one of the most important practical stages of the track: Molecular Docking and Virtual Screening Foundations. This part of the bundle covers the theoretical basis of docking, binding affinity, interaction logic, result analysis, and practical simulation workflows. It also introduces docking applications across different systems, including protein–ligand docking, protein–protein docking, DNA/RNA docking, and even nanomaterial docking, using professional tools such as MOE, Schrödinger, and AutoDock Vina.
The final layer of the track focuses on scientific writing, presentation skills, and collaboration. This is essential because the goal of the track is not only to teach analysis, but to help the learner produce outputs that are more professional, more communicable, and more useful in academic, research, or client-facing environments. By the end of the track, the learner has not only studied important workflows, but has also built a more organized understanding of how to present analytical work clearly and professionally.
What makes this bundle especially valuable is that it is designed around a service mindset. The learner does not finish with scattered knowledge only. They finish with a much clearer answer to questions such as:
- What analyses can I actually perform?
- What kind of scientific output can I deliver?
- What can I offer to a researcher, project owner, or scientific client?
- How can I start building a profile around these skills?
That is why this Service Track also includes a Science Freelance project-oriented layer. This is not a full research project like the one found in larger flagship programs. Instead, it is a service-oriented project layer designed to help the learner organize their work professionally, understand their deliverables, build a stronger analysis identity, and begin structuring their technical output within Science Freelance. This creates a bridge between learning and practical professional positioning.
In short, Service Track 01 is ideal for learners who want a focused, practical, and affordable pathway into bioinformatics, protein-centered analysis, and molecular docking services, without needing to enter the full-scale FAB Program from the beginning.
What You Will Learn
By the end of this Service Track, you will be able to:
- understand the scientific foundation of Chemoinformatics and Computational Drug Discovery
- explain the logic of Computer-Aided Drug Design (CADD) workflows
- navigate major chemical and protein databases
- work with important molecular file formats used in CADD workflows
- understand protein structure, function, folding, and misfolding concepts
- perform protein structure prediction workflows
- apply homology modeling principles and tools
- understand ab initio modeling concepts
- visualize protein structures and assess structural features
- evaluate protein models and understand validation logic
- perform pairwise sequence alignment
- perform multiple sequence alignment (MSA)
- construct and interpret phylogenetic trees
- analyze conserved sequence regions and sequence–function relationships
- perform gene prediction workflows
- analyze protein–protein interaction (PPI) networks
- understand the theory and workflow of molecular docking
- perform docking-oriented workflows for multiple biological systems
- understand virtual screening foundations
- analyze docking outputs and interaction-related results
- produce clearer scientific reports and structured outputs
- build a practical and professional analysis/service list
- begin organizing your technical work through Science Freelance
List of Analyses You Will Be Able to Perform
One of the most important outcomes of this Service Track is that the learner finishes with a clear understanding of the analyses and services they can perform professionally.
Core Analysis and Service List
1. Database Search and Scientific Data Retrieval
Retrieve relevant chemical, protein, and biological data from major scientific databases such as PubChem, DrugBank, PDB, UniProt, KEGG, ZINC, ChEMBL, and related platforms.
2. Molecular File Preparation and Format Handling
Work with key molecular and structural file formats such as SDF, MOL, PDB, and other research-relevant formats used in CADD and bioinformatics pipelines.
3. Protein Sequence Analysis
Analyze protein sequences to identify patterns, features, and relationships relevant to structure, function, and downstream workflows.
4. Pairwise Sequence Alignment
Compare two biological sequences to study similarity, divergence, and possible functional relationships.
5. Multiple Sequence Alignment (MSA)
Align multiple sequences to detect conserved regions and support comparative biological interpretation.
6. Phylogenetic Tree Construction and Interpretation
Build and interpret phylogenetic trees to study evolutionary relationships and sequence-based biological insights.
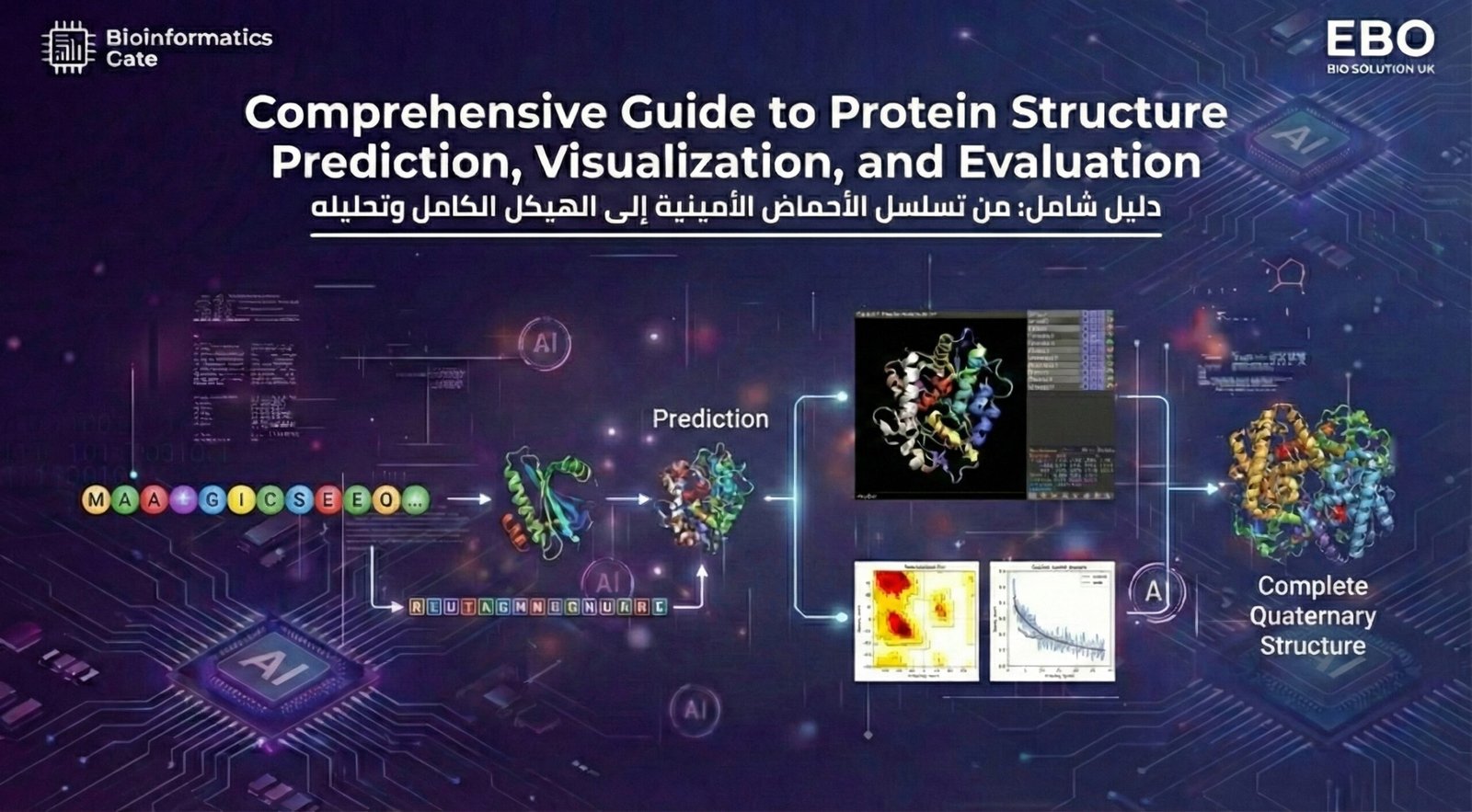
7. Protein Structure Prediction
Predict protein structures using computational tools and structured workflows relevant to bioinformatics and molecular modeling.
8. Homology Modeling
Build 3D protein models based on structurally related templates using homology modeling methodologies.
9. Protein Structure Visualization
Visualize and inspect protein structures using molecular modeling and structural biology tools.
10. Protein Model Evaluation and Validation
Assess model quality, structural acceptability, and biological relevance using validation and quality assessment logic.
11. Gene Prediction
Use bioinformatics tools and genomic sequence logic to identify and annotate genes computationally.
12. Protein–Protein Interaction (PPI) Analysis
Analyze protein interaction networks and interpret their biological and pathway-level significance.
13. Target-Oriented Protein Analysis
Prepare and analyze protein targets in a way that supports docking, interaction studies, and downstream computational workflows.
14. Molecular Docking – Protein–Ligand
Perform docking studies between ligands and protein targets and interpret binding-related results.
15. Molecular Docking – Protein–Protein
Apply docking workflows to protein–protein systems when relevant to research questions.
16. Molecular Docking – DNA / RNA Systems
Understand and perform docking workflows involving DNA or RNA systems when required.
17. Nanomaterial Docking Foundations
Apply docking principles to nanomaterial–biological target systems in relevant cases.
18. Virtual Screening Foundations
Understand and perform the core steps of virtual screening workflows for identifying candidate molecules.
19. Docking Result Analysis and Interaction Interpretation
Analyze docking poses, interaction patterns, and binding-related outputs in a clearer scientific way.
20. Scientific Report Writing
Organize analysis outputs into clearer scientific reports and structured written deliverables.
21. Presentation-Ready Scientific Output
Prepare analysis results in a format suitable for academic presentation, reporting, or professional communication.
What This Analysis List Means
This analysis list means that the learner does not leave the track with general theory only. They leave with a much clearer professional map of what they can actually do.
This includes five major output layers:
1. Data and Database Work
The ability to retrieve, organize, and prepare scientific data correctly.
2. Protein-Centered Analysis
The ability to work with protein sequence, structure, modeling, visualization, and evaluation.
3. Genomics and Interaction Analysis
The ability to perform gene prediction and PPI-related analytical tasks.
4. Docking-Oriented Services
The ability to perform and interpret multiple forms of molecular docking and foundational virtual screening.
5. Scientific Output and Communication
The ability to present results through clearer reports, structured outputs, and professional communication.
Target Audience
This Service Track is ideal for:
- students in pharmacy, biotechnology, bioinformatics, biology, chemistry, and related scientific fields
- recent graduates who want a practical entry point into CADD
- researchers who want to build a stronger computational workflow foundation
- learners who want a smaller and more affordable alternative to a full FAB Program
- early-stage scientific freelancers who want a clearer list of professional analyses
- anyone who wants to begin with bioinformatics, protein workflows, and molecular docking services in a more structured way
Requirements
This Service Track does not require advanced expertise before joining, but it is best suited for participants who have:
- a real interest in bioinformatics, computational drug discovery, or protein-centered analysis
- willingness to follow a structured learning pathway
- readiness to study consistently and apply what they learn
- openness to building practical and professional scientific output
- interest in organizing their work inside Science Freelance
A basic background in one or more of the following is helpful:
- biology
- chemistry
- pharmacy
- biotechnology
- bioinformatics
- computational science
Science Freelance Project Layer
This Service Track includes a Science Freelance project-oriented layer.
This is not a full research project. Instead, it is a structured practical layer that helps the learner begin turning technical learning into a more visible and organized professional output.
Through this layer, the learner begins to:
- organize their analysis identity
- understand how their work can be presented as a service
- structure their outputs more professionally
- connect learning with practical visibility
- start building a stronger profile inside Science Freelance
Optional Add-On A – Client Reach Support Package
This Service Track may also offer Add-On A, an optional monthly support package for participants who want help reaching fresh client opportunities.
Price: 2,300 EGP per month
This add-on may include:
- campaign content preparation
- ad launch and campaign management
- scheduled campaign cycles
- fresh contact lists / new leads generated from active campaigns
- equal distribution among subscribed participants inside the same Service Track
This add-on is optional and is not included in the core Service Track price. It supports audience and client reach, but does not guarantee final purchase or final client conversion.